Experiment Settings¶
This section describes the settings that are available when running an experiment.
Display Name¶
Optional: Specify a display name for the new experiment. There are no character or length restrictions for naming. If this field is left blank, Driverless AI will automatically generate a name for the experiment.
Dropped Columns¶
Dropped columns are columns that you do not want to be used as predictors in the experiment. Note that Driverless AI will automatically drop ID columns and columns that contain a significant number of unique values (above max_relative_cardinality in the config.toml file or Max. allowed fraction of uniques for integer and categorical cols in Expert settings).
Validation Dataset¶
The validation dataset is used for tuning the modeling pipeline. If provided, the entire training data will be used for training, and validation of the modeling pipeline is performed with only this validation dataset. When you do not include a validation dataset, Driverless AI will do K-fold cross validation for I.I.D. experiments and multiple rolling window validation splits for time series experiments. For this reason it is not generally recommended to include a validation dataset as you are then validating on only a single dataset. Note that time series experiments cannot be used with a validation dataset: including a validation dataset will disable the ability to select a time column and vice versa.
This dataset must have the same number of columns (and column types) as the training dataset. Also note that if provided, the validation set is not sampled down, so it can lead to large memory usage, even if accuracy=1 (which reduces the train size).
Test Dataset¶
The test dataset is used for testing the modeling pipeline and creating test predictions. The test set is never used during training of the modeling pipeline. (Results are the same whether a test set is provided or not.) If a test dataset is provided, then test set predictions will be available at the end of the experiment.
Weight Column¶
Optional: Column that indicates the observation weight (a.k.a. sample or row weight), if applicable. This column must be numeric with values >= 0. Rows with higher weights have higher importance. The weight affects model training through a weighted loss function and affects model scoring through weighted metrics. The weight column is not used when making test set predictions, but a weight column (if specified) is used when computing the test score.
Note: The weight column is not used as a feature in modeling.
Fold Column¶
Optional: Rows with the same value in the fold column represent groups that should be kept together in the training, validation, or cross-validation datasets. This can prevent data leakage and improve generalization for data that is naturally grouped and not i.i.d. (identically and independently distributed). This column must be an integer or categorical variable, and it cannot be specified if a validation set is used or if a Time Column is specified.
By default, Driverless AI assumes that the dataset is i.i.d. and creates validation datasets randomly for regression or with stratification of the target variable for classification.
The fold column is used to create the training and validation datasets so that all rows with the same Fold value will be in the same dataset. This can prevent data leakage and improve generalization. For example, when viewing data for a pneumonia dataset, person_id would be a good Fold Column. This is because the data may include multiple diagnostic snapshots per person, and we want to ensure that the same person’s characteristics show up only in either the training or validation frames, but not in both to avoid data leakage.
This column must be an integer or categorical variable and cannot be specified if a validation set is used or if a Time Column is specified.
Note: The fold column is not used as a feature in modeling.
Time Column¶
Optional: Specify a column that provides a time order (time stamps for observations), if applicable. This can improve model performance and model validation accuracy for problems where the target values are auto-correlated with respect to the ordering (per time-series group).
The values in this column must be a datetime format understood by pandas.to_datetime(), like “2017-11-29 00:30:35” or “2017/11/29”, or integer values. If [AUTO] is selected, all string columns are tested for potential date/datetime content and considered as potential time columns. If a time column is found, feature engineering and model validation will respect the causality of time. If [OFF] is selected, no time order is used for modeling and data may be shuffled randomly (any potential temporal causality will be ignored).
When your data has a date column, then in most cases, specifying [AUTO] for the Time Column will be sufficient. However, if you select a specific date column, then Driverless AI will provide you with an additional side menu. From this side menu, you can specify Time Group columns or specify [Auto] to let Driverless AI determine the best time group columns. You can also specify the columns that will be unavailable at prediction time (see More About Unavailable Columns at Time of Prediction for more information), the Forecast Horizon (in a unit of time identified by Driverless AI), and the Gap between the train and test periods.
Refer to Time Series in Driverless AI for more information about time series experiments in Driverless AI and to see a time series example.
Notes:
Engineered features will be used for MLI when a time series experiment is built. This is because munged time series features are more useful features for MLI compared to raw time series features.
A Time Column cannot be specified if a Fold Column is specified. This is because both fold and time columns are only used to split training datasets into training/validation, so once you split by time, you cannot also split with the fold column. If a Time Column is specified, then the time group columns play the role of the fold column for time series.
A Time Column cannot be specified if a validation dataset is used.
A column that is specified as being unavailable at prediction time will only have lag-related features created for (or with) it.
Unavailable Columns at Time of Prediction will only have lag-related features created for (or with) it, so this option is only used when time-series-lag-based-recipe is enabled.
Accuracy, Time, and Interpretability Knobs¶
The experiment preview describes what the Accuracy, Time, and Interpretability settings mean for your specific experiment. This preview automatically updates when any of the experiment’s settings change (including the knobs).
The Accuracy knob stands for relative accuracy: Higher values should lead to higher confidence in model performance. Usually achieved through the use of larger data (less sampling), more modeling effort (more tuning, higher accuracy settings), more statistical calculations (cross-validation, bootstrapping). Doesn’t always mean that the final model is better, but generally means that the final estimate is more accurate. If in doubt, trust the results of the experiment with higher accuracy settings.
The Time knob stands for relative time tolerance: Higher values generally lead to longer run times. Indicates patience to wait for convergence of the experiment score. Larger values mean higher chance of getting a better model. If it takes too long, just click on ‘Finish’ button and it will finish the experiment as if convergence was achieved.
The Interpretability knob stands for relative interpretability: Higher values favor more interpretable models (e.g. linear models, decision trees, single models) with less complex feature engineering (fewer features, simple features). Lower values lead to more complex models (e.g. neural networks, GBMs, ensembles) and more complex feature pipelines (more features, higher-order interaction features).
Note
You can manually select individual features to force into an experiment—regardless of Accuracy, Time, and Interpretability levels—with the Features to Force In expert setting.
To adjust the lowest allowed variable importance that features can have before being dropped, use the Lowest Allowed Variable Importance at Interpretability 10 expert setting.
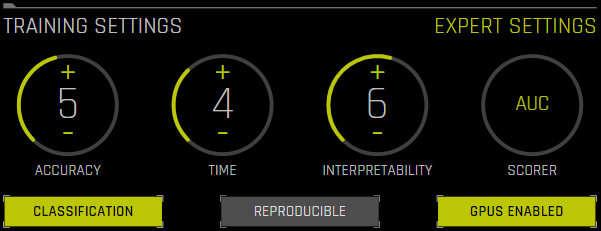
Accuracy, Time, and Interpretability Knobs¶

Experiment Preview¶
Accuracy¶
As accuracy increases, Driverless AI gradually adjusts the method for performing the evolution and ensemble. At low accuracy, Driverless AI varies features and models, but they all compete evenly against each other. At higher accuracy, each independent main model will evolve independently and be part of the final ensemble as an ensemble over different main models. At higher accuracies, Driverless AI will evolve+ensemble feature types like Target Encoding on and off that evolve independently. Finally, at highest accuracies, Driverless AI performs both model and feature tracking and ensembles all those variations.
Changing this value affects the feature evolution and final pipeline.
Note: A check for a shift in the distribution between train and test is done for accuracy >= 5.
Training data size: Displays the number of rows and columns in the training data.
Feature evolution: This represents the algorithms used to create the experiment. If a test set is provided without a validation set, then Driverless AI will perform a 1/3 validation split during the experiment. If a validation set is provided, then the experiment will perform external validation.
Final pipeline: This represents the number of models and the validation method used in the final pipeline. For ensemble modeling, information about how models are combined is also shown here.
Time¶
This specifies the relative time for completing the experiment (that is, higher settings take longer).
Feature Brain Level: Displays the feature brain level for the experiment. For more information, see feature_brain1.
Feature evolution: Displays the number of individuals and maximum number of iterations that will be run in this experiment.
Early stopping: Early stopping will take place if the experiment doesn’t improve the score for the specified amount of iterations.
Interpretability¶
Specify the relative interpretability for this experiment. Higher values favor more interpretable models. Changing the interpretability level affects the feature pre-pruning strategy, monotonicity constraints, and the feature engineering search space.
Feature pre-pruning strategy: This represents the feature selection strategy (to prune-away features that do not clearly give improvement to model score). Strategy = “Permutation Importance FS” if interpretability >= 6; otherwise strategy is None.
Monotonicity constraints: If Monotonicity Constraints are enabled, the model will satisfy knowledge about monotonicity in the data and monotone relationships between the predictors and the target variable. For example, in house price prediction, the house price should increase with lot size and number of rooms, and should decrease with crime rate in the area. If enabled, Driverless AI will automatically determine if monotonicity is present and enforce it in its modeling pipelines. Depending on the correlation, Driverless AI will assign positive, negative, or no monotonicity constraints. Monotonicity is enforced if the absolute correlation is greater than 0.1. All other predictors will not have monotonicity enforced. For more information, see Monotonicity Constraints.
Note: Monotonicity constraints are used in XGBoost GBM, XGBoost Dart, LightGBM, and Decision Tree models.
Feature engineering search space: This represents the transformers that will be used during the experiment.
[…] Models to Train¶
For the listed models:
Model and feature tuning: Represents the number of validation splits multiplied by the tuning population size.
Feature evolution: Represents the number of models trained in order to evaluate engineered features.
Final pipeline: Represents the number of final models.
Per-model hyperparameter optimization trials:
evolution - Represents the number of trials performed for hyperparameter optimization for tuning models.
final - Represents the number of trials performed for hyperparameter optimization for final models.
Classification/Regression Button¶
Driverless AI automatically determines the problem type based on the response column. Though not recommended, you can override this setting by clicking this button.
Reproducible¶
The Reproducible toggle lets you build an experiment with a random seed and get reproducible results. If this is disabled (default), then results vary between runs, which can give a good sense of variance among experiment results.
When enabling this option, keep the following notes in mind:
Experiments are only reproducible when run on the same hardware (that is, using the same number and type of GPUs/CPUs and the same architecture). For example, you will not get the same results if you try an experiment on a GPU machine, and then attempt to reproduce the results on a CPU-only machine or on a machine with a different number and type of GPUs.
This option should be used with the reproducibility_level expert setting option, which ensures different degrees of reproducibility based on the OS and environment architecture. Keep in mind that when Reproducibility is enabled, then
reproducibility_level=1by default.Experiments run using TensorFlow with multiple cores cannot be reproduced.
LightGBM is more reproducible with 64-bit floats, and Driverless AI will switch to 64-bit floats for LightGBM. (Refer to https://lightgbm.readthedocs.io/en/latest/Parameters.html#gpu_use_dp for more information.)
Enabling this option automatically disables all of the Feature Brain expert settings options; specifically:
feature_brain1
feature_brain2
feature_brain3
feature_brain4
feature_brain5
For more information on reproducibility in Driverless AI, see Reproducibility in Driverless AI.
Enable GPUs¶
You can enable GPU support when setting up an experiment by clicking the GPU On or Off buttons.
Notes:
Only supported on machines with NVIDIA GPU accelerators. For best results, we recommend GPUs that use the Pascal or Volta architectures.
Image and natural language processing (NLP) use cases in H2O Driverless AI benefit significantly from GPU usage.
This option is ignored on CPU-only systems.
